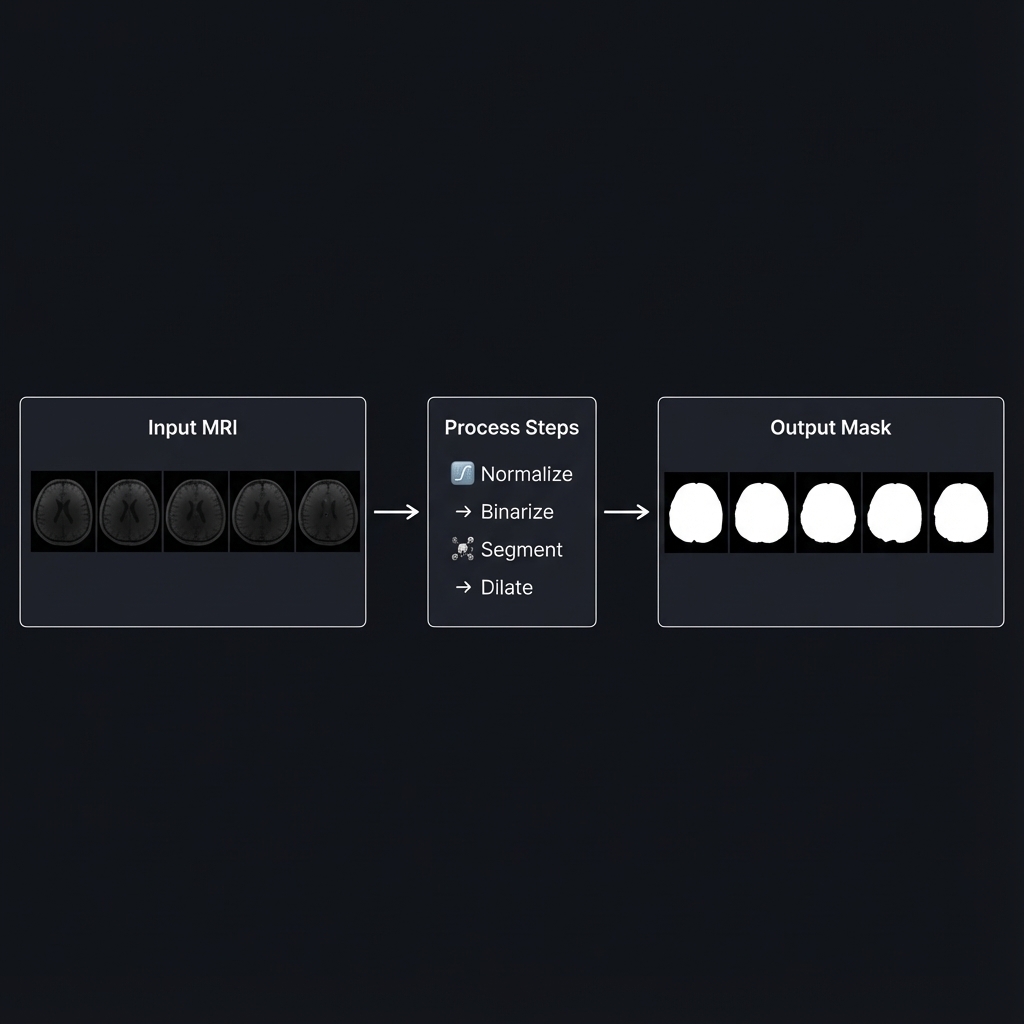
Building a Skull Mask Generator for MRI Images Without Deep Learning
A classical image processing approach to skull mask generation, achieving IoU 0.98 with only 5 labeled images—no deep learning required.
Introduction
Recently, I worked on a task to generate skull masks from MRI images. The goal was simple:
- Exclude the outermost bright tissue (scalp)
- Include the dark border inside (skull bone)
My first instinct was U-Net. But I only had 5 labeled images—not enough to train anything.
So I went with classical image processing instead. The result: IoU 0.9795, Dice 0.9896.
The Data
The dataset consisted of:
- Input: 5 MRI slices, shape
(5, 768, 624), dtypefloat32 - Ground Truth: 5 binary masks, shape
(5, 768, 624), dtypeuint8
The first thing I noticed was the unusual value range of the input images:
1
2
Min: 1.83e-07
Max: 3.49e-05
These are extremely small values—not the typical 0-255 range you’d expect. This immediately told me that normalization would be essential before applying any standard image processing operations.
Algorithm Overview
After several iterations (more on that later), I settled on this pipeline:
flowchart LR
subgraph Input
A[MRI Image<br/>float32]
end
subgraph Processing
B[Normalize<br/>0-255] --> C[Otsu<br/>Binarization]
C --> D[Connected<br/>Components]
D --> E{Edge<br/>Touching?}
E -->|Yes| F[Scalp]
E -->|No| G[Brain]
G --> H[Dilation<br/>26px]
H --> I[Remove<br/>Scalp]
I --> J[Fill<br/>Holes]
end
subgraph Output
K[Skull Mask<br/>uint8]
end
A --> B
J --> K
style F fill:#ff6b6b,color:#fff
style G fill:#51cf66,color:#fff
style K fill:#339af0,color:#fff
The key insight: Scalp touches the image border, Brain does not.
Step 1: Normalization
Why Normalize?
OpenCV functions expect pixel values in the 0-255 range (uint8). Our raw MRI data ranges from 1.8e-07 to 3.5e-05. If we feed these values directly to OpenCV, the functions won’t work as expected.
How It Works
1
2
3
4
5
6
7
def normalize_image(image: np.ndarray) -> np.ndarray:
img_min = image.min()
img_max = image.max()
img_norm = (image - img_min) / (img_max - img_min)
img_uint8 = (img_norm * 255).astype(np.uint8)
return img_uint8
# end def
Before: Values in range [1.8e-07, 3.5e-05] After: Values in range [0, 255]
Step 2: Otsu Binarization
What Is Binarization?
Binarization converts a grayscale image into a binary image (black and white). Every pixel becomes either 0 (black) or 1 (white) based on a threshold.
The Threshold Selection Problem
If we choose threshold = 100:
- Pixels ≥ 100 → White (1)
- Pixels < 100 → Black (0)
But how do we know 100 is the right value? What if 80 or 120 is better?
Otsu’s Algorithm: Automatic Threshold Selection
Otsu’s method analyzes the histogram and automatically finds the optimal threshold that best separates the two classes (foreground and background).
1
_, binary = cv2.threshold(img_uint8, 0, 255, cv2.THRESH_BINARY + cv2.THRESH_OTSU)
For our MRI images, Otsu consistently found threshold = 23:
1
2
3
4
5
6
7
8
9
10
Histogram Distribution:
Count
│
│ ████ ████
│ ████ ████
│──────────────────────────────────── Brightness
0 23 200
↑
Otsu's optimal split point
Interpretation: Pixels with brightness < 23 are background/skull bone (dark), and pixels ≥ 23 are brain/scalp tissue (bright).
Step 3: Connected Component Analysis
The Key Insight
After binarization, we have bright regions (tissue) and dark regions (background). But how do we distinguish between Scalp (which we want to remove) and Brain (which we want to keep)?
Here’s the crucial observation:
| Region | Characteristic |
|---|---|
| Scalp | Touches the image border |
| Brain | Does NOT touch the image border |
This makes sense anatomically: the scalp wraps around the head and extends to the edges of the MRI slice, while the brain is enclosed inside.
Visual Explanation
1
2
3
4
5
6
7
8
9
10
11
12
13
14
MRI Image (Binarized):
┌─────────────────────────────────┐
│█████████████████████████████████│← Touches top edge (Scalp)
│██ ██│
│█ █│← Touches left/right edges (Scalp)
│█ ┌───────────────────┐ █│
│█ │ │ █│
│█ │ Brain │ █│← No edge contact (Brain)
│█ │ │ █│
│█ └───────────────────┘ █│
│██ ██│
│█████████████████████████████████│← Touches bottom edge (Scalp)
└─────────────────────────────────┘
Implementation
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
for label in range(1, num_labels):
component = (labels == label)
touches_edge = (
np.any(component[0, :]) or # Top row
np.any(component[-1, :]) or # Bottom row
np.any(component[:, 0]) or # Left column
np.any(component[:, -1]) # Right column
)
if touches_edge:
# This is Scalp → Remove later
edge_labels.add(label)
else:
# This is Brain → Keep
inner_labels.append(label)
# end if
# end for
Step 4: Morphological Dilation
The Problem
After identifying the brain region, we’re not done yet. The task requires us to include the dark border (skull bone), not just the brain tissue.
The anatomical structure from outside to inside:
1
2
3
[Scalp] → [Skull Bone] → [Brain]
Bright Dark Bright
Remove Include Include
If we only keep the brain region, we miss the skull bone entirely.
The Solution: Dilation
Dilation expands a region by a certain number of pixels in all directions.
1
2
3
4
5
6
7
8
Before Dilation: After Dilation (26px):
┌─────────┐ ┌───────────────┐
│ │ │███████████████│
│ Brain │ → │███ Brain ███│
│ │ │███████████████│
└─────────┘ └───────────────┘
The expanded area covers the Skull Bone region!
Why 26 Pixels?
I tested various dilation sizes:
| Dilation Size | IoU | Dice |
|---|---|---|
| 22 | 0.9742 | 0.9869 |
| 24 | 0.9776 | 0.9887 |
| 26 | 0.9794 | 0.9896 |
| 28 | 0.9788 | 0.9893 |
| 30 | 0.9763 | 0.9880 |
At 26 pixels, the expanded brain region perfectly captures the skull bone while not extending too far into the scalp.
Step 5: Hole Filling
Why Fill Holes?
After dilation, there might be small holes inside the mask—caused by dark structures within the brain (like ventricles) that were classified as background during binarization.
1
2
3
4
5
6
7
8
9
10
Before Filling: After Filling:
┌─────────────┐ ┌─────────────┐
│█████████████│ │█████████████│
│███ █████│ │█████████████│
│███ ○ █████│ → │█████████████│
│███ █████│ │█████████████│
│█████████████│ │█████████████│
└─────────────┘ └─────────────┘
↑
Internal hole
The skull mask should include the entire interior of the skull, so we fill any internal holes:
1
skull_mask = ndimage.binary_fill_holes(skull_mask).astype(np.uint8)
Understanding the Metrics
IoU (Intersection over Union)
\[\text{IoU} = \frac{\text{Predicted} \cap \text{Ground Truth}}{\text{Predicted} \cup \text{Ground Truth}}\]- IoU = 1.0: Perfect overlap
- IoU = 0.0: No overlap at all
Our result: IoU = 0.9794 means 97.94% overlap with the ground truth.
Dice Coefficient
\[\text{Dice} = \frac{2 \times |\text{Predicted} \cap \text{Ground Truth}|}{|\text{Predicted}| + |\text{Ground Truth}|}\]Our result: Dice = 0.9896 is considered excellent in medical image segmentation.
The Development Journey
I didn’t arrive at this solution immediately. Here’s my iteration history:
| Version | Approach | IoU | Issue |
|---|---|---|---|
| v1 | Otsu + Largest Component | 0.73 | Skull bone not included |
| v2 | Flood Fill from Edges | 0.82 | Incomplete scalp removal |
| v3-v4 | Various Thresholds | 0.54 | Made it worse |
| v5 | Connected Components + Dilation | 0.96 | Key breakthrough |
| v6 | Dilation Size Tuning | 0.97 | Improved |
| v7 | Optimized Dilation (26px) | 0.98 | Final |
The key insight came in v5 when I realized that edge-touching was the discriminating feature between scalp and brain, not just brightness values.
Why Not Deep Learning?
Given that this is a segmentation task, you might wonder why I didn’t use U-Net or a similar architecture. Here’s my reasoning:
| Criterion | Classical Approach | Deep Learning |
|---|---|---|
| Training Data | Not needed | 500-1,000+ images minimum |
| 2D Slice Support | ✅ | ✅ |
| External Dependencies | numpy, opencv, scipy | GPU, large frameworks |
| Interpretability | Each step is clear | Black box |
| Current Accuracy | IoU 0.98 | Potentially higher with enough data |
With only 5 labeled images, deep learning was simply not an option. However, if 10,000+ labeled images became available, I would definitely consider training a U-Net for potentially even better results.
Scaling to 100,000 Images
As part of the project, I also planned for processing 100,000 images within 2 weeks.
Performance Analysis
| Metric | Value |
|---|---|
| Processing time per image | 16.6 ms |
| Images per second | 60.2 |
| Time for 100,000 images (single thread) | ~28 minutes |
| Time for 100,000 images (8 cores) | ~4 minutes |
The classical approach turned out to be extremely fast—no GPU required!
Batch Processing Script
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
import numpy as np
from multiprocessing import Pool
from skull_mask_generator import create_skull_mask
def process_single(image: np.ndarray) -> np.ndarray:
return create_skull_mask(image)
# end def
if __name__ == '__main__':
images = np.load('large_dataset.npy')
with Pool(processes=8) as pool:
masks = pool.map(process_single, images)
# end with
np.save('output_masks.npy', np.array(masks))
# end if
Conclusion
Key insights:
- Edge-touching property was the breakthrough—Scalp touches borders, Brain doesn’t
- Iterate and measure—v1-v4 failed, v5 worked
- Simple can be powerful—Otsu + Connected Components + Dilation = 98% IoU
Not every problem needs deep learning.